Sanders, R. N. C., Jones, D. C., Josey, S. A., Sinha, B., and Forget, G. (2022), Causes of the 2015 North Atlantic cold anomaly in a global state estimate, Ocean Sci., doi: 10.5194/os-18-953-2022 Continue reading Causes of the 2015 North Atlantic cold anomaly in a global state estimate
Tag Archives: Forget
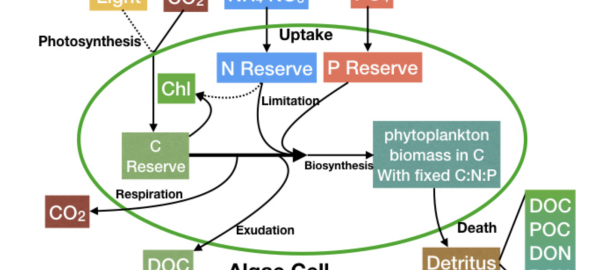
A Super-Speedy New Phytoplankton Model
Darwin Group researchers Zhen Wu and Gael Forget apply Julia to create a GPU-supported individual-based phytoplankton life cycle model. Continue reading A Super-Speedy New Phytoplankton Model
Christopher L. Follett, Stephanie Dutkiewicz, Gael Forget, B.B. Cael, Michael J. Follows (2021), Moving ecological and biogeochemical transitions across the North Pacific, Limnology and Oceanography, doi: 10.1002/lno.11763 Continue reading Moving ecological and biogeochemical transitions across the North Pacific
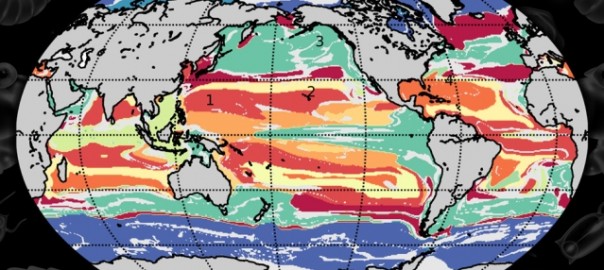
Machine learning helps map global ocean communities
Technique developed by MIT-CBIOMES investigators could aid in tracking the ocean’s health and productivity. Continue reading Machine learning helps map global ocean communities
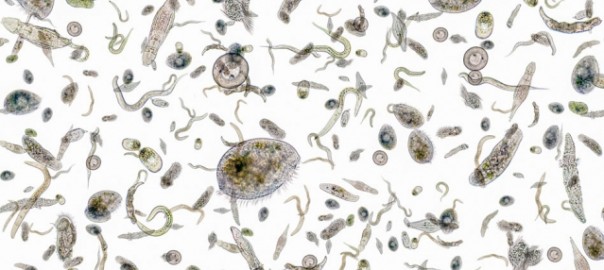
Seeding Oceans with Iron may not Impact Climate Change
Darwin Project study led by Jonathan Lauderdale finds Earth’s oceans contain just the right amount of iron; adding more may not improve their ability to absorb carbon dioxide. Continue reading Seeding Oceans with Iron may not Impact Climate Change

Darwin Goes to Ocean Sciences 2020
Look out for the Darwin team, sharing their work at this year’s Ocean Sciences conference taking place February 16-21 in San Diego, California.
Dean Roemmich, Matthew H. Alford, Hervé Claustre, Kenneth Johnson, Brian King, James Moum, Peter Oke, W. Brechner Owens, Sylvie Pouliquen, Sarah Purkey, Megan Scanderbeg, Toshio Suga, Susan Wijffels, Nathalie Zilberman, Dorothee Bakker, Molly Baringer, Mathieu Belbeoch, Henry C. Bittig, Emmanuel Boss, Paulo Calil, Fiona Carse, Thierry Carval, Fei Chai, Diarmuid Ó. Conchubhair, Fabrizio d’Ortenzio, Giorgio Dall’Olmo, Damien Desbruyeres, Katja Fennel, Ilker Fer, Raffaele Ferrari, Gaël Forget, Howard Freeland, Tetsuichi Fujiki, Marion Gehlen, Blair Greenan, Robert Hallberg, Toshiyuki Hibiya, Shigeki Hosoda, Steven Jayne, Markus Jochum, Gregory C. Johnson, KiRyong Kang, Nicolas Kolodziejczyk, Arne Körtzinger, Pierre-Yves Le Traon, Yueng-Djern Lenn, Guillaume Maze, Kjell Arne Mork, Tamaryn Morris, Takeyoshi Nagai, Jonathan Nash, Alberto Naveira Garabato, Are Olsen, Rama Rao Pattabhi, Satya Prakash, Stephen Riser, Catherine Schmechtig, Claudia Schmid, Emily Shroyer, Andreas Sterl, Philip Sutton, Lynne Talley, Toste Tanhua, Virginie Thierry, Sandy Thomalla, John Toole, Ariel Troisi, Thomas W. Trull, Jon Turton, Pedro Joaquin Velez-Belchi, Waldemar Walczowski, Haili Wang, Rik Wanninkhof, Amy F. Waterhouse, Stephanie Waterman, Andrew Watson, Cara Wilson, Annie P. S. Wong, Jianping Xu and Ichiro Yasuda (2019), On the Future of Argo: A Global, Full-Depth, Multi-Disciplinary Array, Frontiers of Marine Science, doi: 10.3389/fmars.2019.00439 Continue reading On the Future of Argo: A Global, Full-Depth, Multi-Disciplinary Array
Andrea Storto, Aida Alvera-Azcárate, Magdalena A. Balmaseda, Alexander Barth, Matthieu Chevallier, Francois Counillon, Catia M. Domingues, Marie Drevillon, Yann Drillet, Gaël Forget, Gilles Garric, Keith Haines, Fabrice Hernandez, Doroteaciro Iovino, Laura C. Jackson, Jean-Michel Lellouche, Simona Masina, Michael Mayer, Peter R. Oke, Stephen G. Penny, K. Andrew Peterson, Chunxue Yang and Hao Zuo (2019), Ocean Reanalyses: Recent Advances and Unsolved Challenges, Frontiers of Marine Science, doi: 10.3389/fmars.2019.00418 Continue reading Ocean Reanalyses: Recent Advances and Unsolved Challenges